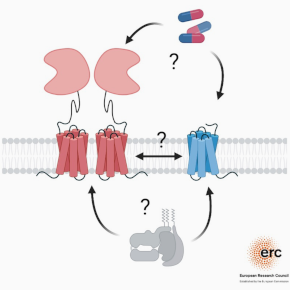
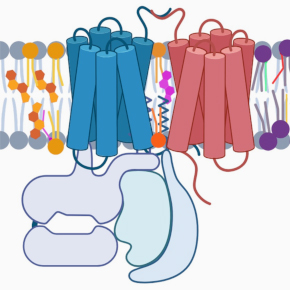
mGlu and GABA-B receptors are complex G protein-coupled receptors that function as homodimers or heterodimers. They are also capable of forming complexes with other proteins, including membrane proteins, to modulate their activity in vivo. It is important to better understand the molecular and structural basis of these complexes and their dynamics, in particular to develop new molecules of therapeutic interest in the field of neurological and psychiatric diseases.
The team’s research project has four specific objectives:
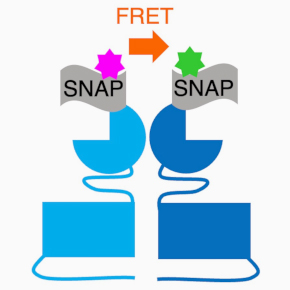
1 – To study the molecular and structural basis of mGlu receptors and the complexes they form with other membrane partners, using biophysical and structural approaches (cryo-electron microscopy, analysis of conformational dynamics at the single-molecule scale using the Förster-type resonance energy transfer (FRET) techniques).
2- To develop methods and tools for studying mGlu and GABA-B receptors in vitro and in their native environment. The preferred methods are biosensors based on Förster-type resonance energy transfer (FRET, HTRFÒ) or bioluminescence (BRET) technologies. The tools developed are specific ligands, small molecules or camelid antibodies (nanobodies), and some of them can be controlled by light.
3- To study the complexes formed by mGlu receptors and their dynamics in their native environment. The aim is to identify functional complexes in the brain, such as the heterodimeric mGlu receptors recently discovered by the team. And to understand the physiological importance of these complexes for neuronal and brain function. The tools described above are used.
4- To develop nanobodies targeting mGlu receptors as innovative immunotherapies for the treatment of brain diseases, in particular schizophrenia.
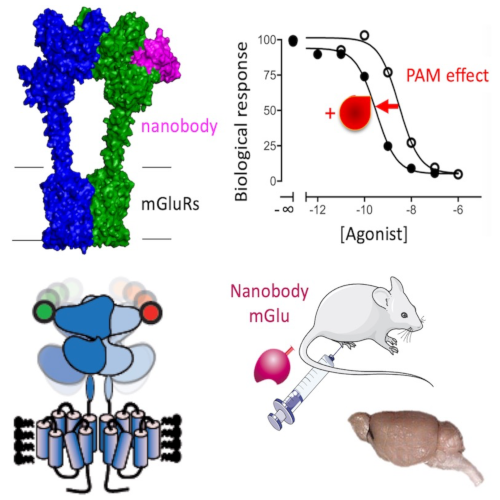
The team studies the molecular pharmacology of mGlu and GABA-B receptors, from their structure and activation mechanism, pharmacology and signaling, to the development of innovative new ligands (camelid nanobodies and photoactivatable compounds) and their applications in several models of brain diseases.
Team leader

DR1, Inserm

IGF Nord 223

04 34 35 92 89
Know more >
Researchers

CRCN, CNRS

IGF Nord 209b

04 34 35 92 78
Know more >

DRCE, CNRS

IGF Nord 223

04 34 35 92 89
Know more >

DR2, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >

Accueil scientifique, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >
Technicians and engineers

IEHC, CNRS

IGF Nord 219

04 34 35 93 13
Know more >

IECN, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >
Postdoctoral researchers and doctoral students

Doctorant(e), UM

IGF Nord 223

04 34 35 93 07
Know more >

Chercheur CDD, CNRS

IGF Nord 215a

04 34 35 93 07
Know more >

Chercheur CDD, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >

Chercheur CDD, CNRS

IGF Nord 223

04 34 35 92 78
Know more >

Doctorant(e), UM

IGF Nord 221a

04 34 35 92 78
Know more >

Chercheur CDD, CNRS

IGF Nord 209b

04 34 35 93 07
Know more >

Chercheur CDD, CNRS

IGF Nord 209b

04 34 35 92 89
Know more >

Chercheur CDD, CNRS

IGF Nord 223

04 34 35 92 89
Know more >

Chercheur CDD, CNRS

IGF Nord 216a

04 34 35 92 63
Know more >

Chercheur CDD, CNRS

IGF Nord 215a

04 34 35 92 78
Know more >

Doctorant(e), Autre

IGF Nord 215b

04 34 35 92 78
Know more >

Chercheur CDD, Inserm

IGF Nord 215b

04 34 35 92 78
Know more >

Doctorant(e), UM

IGF Nord 223

04 34 35 93 07
Know more >

Doctorant(e), CNRS

IGF Nord 215a

04 34 35 93 07
Know more >
Fixed term contract technicians and engineers

IE CDD, CNRS

IGF Nord 221a

04 34 35 92 89
Know more >

IE CDD, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >

IE CDD, CNRS

IGF Nord 216a

04 34 35 92 63
Know more >

IE CDD, CNRS

IGF Nord 223

04 34 35 92 78
Know more >

IE CDD, CNRS

IGF Nord 223

04 34 35 92 78
Know more >

IECN CDD, CNRS

IGF Nord 209b

04 34 35 92 89
Know more >
Trainees

Master 2, CNRS

IGF Nord 215b

04 34 35 92 78
Know more >

Master 2, CNRS

IGF Nord 221a

04 34 35 93 07
Know more >

Stagiaire, CNRS

IGF Nord 223

04 34 35 92 89
Know more >
- Xu C, Zhou Y, Liu Y, Lin L, Liu P, Wang X, Xu Z, Pin JP*, Rondard P*, Liu J* (2024). Specific pharmacological and Gi/o protein coupling properties of native GPCRs in neurons. Nat Comm 2024 15(1) :1990. doi: 10.1038/s41467-024-46177-z. PMID: 38443355 * Co-corresponding author.
- Lecat-Guillet N, Quast RB, Liu H, Moeller TC, Rovira X, Soldevila S, Lamarque L, Trinquet E, Liu J, Pin JP, Rondard P*, Margeat E*. Concerted conformational changes control metabotropic glutamate receptor activity. Sci Adv 2023 9 (22) :eadf1378. doi: 10.1126/sciadv.adf1378. PMID: 37267369 * Co-corresponding author.
- Meng J, Xu C, Lafon PA, Roux S, Mathieu M, Zhou R, Scholler P, Blanc E, Becker JAJ, Le Merrer J, González-Maeso J, Chames P, Liu J*, Pin JP*, Rondard P*. Optical biosensors of native membrane protein complexes reveal a high proportion of mGlu heterodimers in the brain. Nat Chem Biol 2022, 18, 894-903. doi: 10.1038/s41589-022-01050-2. PMID: 35681029 * Co-corresponding author.
- Liu J, Tang H, Xu C, Zhou S, Zhu X, Li Y, Prézeau L, Xu T, Pin JP*, Rondard P*, Ji W*, Liu J* (2022) Biased signaling due to oligomerization of a G protein-coupled receptor. Nat Comm 2022 13, 6365. doi: 10.1038/s41467-022-34056-4. PMID: 36289206 * Co-corresponding author.
- Haubrich J, Font J, Goupil-Lamy A, Scholler P, Nevoltris D, Acher F, Chames P, Rondard P, Prézeau L*, Pin J-P*. A nanobody activating metabotropic glutamate receptor 4 discriminates between homo- and heterodimers. Proc Natl Acad Sci U S A 2021 118 (33):e2105848118. doi: 10.1073/pnas.2105848118. PMID: 34385321* Co-corresponding author.
- Cao AM, Quast RB, Fatemi F, Rondard P, Pin JP*, Margeat E* (2021) Allosteric modulators enhance agonist efficacy by increasing the residence time of a GPCR in the active state. Nat Comm 2021 12 (1):5426. doi: 10.1038/s41467-021-25620-5. PMID: 34521824* Co-corresponding author.
- Liu H, Yi P, Zhao W, Wu Y, Acher F, Pin JP*, Liu J*, Rondard P. Illuminating the allosteric modulation of the calcium sensing receptor. Proc Natl Acad Sci U S A 2020 117 (35) 21711-21722. doi: 10.1073/pnas.1922231117. PMID: 32817431 * Co-corresponding author.
- Xue L, Sun Q, Zhao H, Rovira X, Gai S, He Q, Pin JP*, Liu J*, Rondard P. Rearrangement of the transmembrane domain interfaces associated with the activation of a GPCR hetero-oligomer. Nat Commun 2019 10, 2765. doi: 10.1038/s41467-019-10834-5. PMID: 31235691 * Co-corresponding author.
- Scholler P, Nevoltris D, De Blundel D, Bossi S, Moreno D, Rovira X, Møller TC, El-Moustaine D, Mathieu M, Blanc E, McLean H, Dupuis E, Mathis G, Trinquet E, Daniel H, Valjent E, Baty D, Chames P, Rondard P*, Pin JP*. Allosteric nanobodies uncover a role of hippocampal mGlu2 receptor homodimers in contextual fear consolidation. Nat Commun 2017 8 (1):1967. doi: 10.1038/s41467-017-01489-1. PMID: 29213077 * Co-corresponding author.
- Scholler P, Moreno D, Lecat-Guillet N, Doumazane E, Monnier C, Charrier-Savournin F, Fabre L, Chouvet C, Soldevila S, Lamarque L, Donsimoni G, Roux T, Zwier JM, Trinquet E, Rondard P*, Pin JP*. HTS-compatible FRET-based conformational sensors clarify membrane receptor activation. Nat Chem Biol 2017 13, 372-380. doi: 10.1038/nchembio.2286. PMID: 28135236 * Co-corresponding author

Development of methods and tools for studying GPCRs
Principal investigator
Jean-Philippe PIN
Find out more
Complexes of mGlu receptors and their dynamics in native environment
Principal investigator
Laurent PREZEAU
Find out more
Nanobodies targeting mGlu receptors as innovative immunotherapy for the treatment of brain diseases
Principal investigator
Philippe RONDARD
Find out more
Revvity Cisbio
Codolet, France
Emmanuel Margeat
Centre de Biologie Structurale (CBS), Montpellier, France
Francine Acher
Université Paris Cité, Paris, France
Patrick Chames
Aix-Marseille Université, France
Jianfeng Liu
Huazhong University of Science and Technology (HUST), Wuhan, China
Amadeu Llebaria
University of Barcelona, Barcelona, Spain

PhD students
• Alexandre Bouyssou (Genentech, USA)
• Laetitia Comps-Agrar (Genentech, USA)
• Mélanie Da Silva
• Damien Maurel
• Carine Monnier
• Mathieu Oosterlaken
• Marie-Laure Rives
• Pauline Scholler (Revvity)
Post-doctoral fellows
• Fanny Dubois
• Driss El Moustaine
• Nathalie Lecat-Guillet (Revvity)
• Oualid Sbai
• Li Xue (Shanghai Jiao Tong University School of Medicine, Shanghai, China)
Engineers
• Emilie Blanc
• Michaël Mathieu (Revvity)
• Jessica Monnic (Mabqi)
• Roxane Perrin
• Salomé Roux



 IGF Nord 223
IGF Nord 223 04 34 35 92 89
04 34 35 92 89
 IGF Nord 209b
IGF Nord 209b 04 34 35 92 78
04 34 35 92 78
 IGF Nord 223
IGF Nord 223 04 34 35 92 89
04 34 35 92 89
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 219
IGF Nord 219 04 34 35 93 13
04 34 35 93 13
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 223
IGF Nord 223 04 34 35 93 07
04 34 35 93 07
 IGF Nord 215a
IGF Nord 215a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 223
IGF Nord 223 04 34 35 92 78
04 34 35 92 78
 IGF Nord 221a
IGF Nord 221a 04 34 35 92 78
04 34 35 92 78
 IGF Nord 209b
IGF Nord 209b 04 34 35 93 07
04 34 35 93 07
 IGF Nord 209b
IGF Nord 209b 04 34 35 92 89
04 34 35 92 89
 IGF Nord 223
IGF Nord 223 04 34 35 92 89
04 34 35 92 89
 IGF Nord 216a
IGF Nord 216a 04 34 35 92 63
04 34 35 92 63
 IGF Nord 215a
IGF Nord 215a 04 34 35 92 78
04 34 35 92 78
 IGF Nord 215b
IGF Nord 215b 04 34 35 92 78
04 34 35 92 78
 IGF Nord 215b
IGF Nord 215b 04 34 35 92 78
04 34 35 92 78
 IGF Nord 223
IGF Nord 223 04 34 35 93 07
04 34 35 93 07
 IGF Nord 215a
IGF Nord 215a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 221a
IGF Nord 221a 04 34 35 92 89
04 34 35 92 89
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 216a
IGF Nord 216a 04 34 35 92 63
04 34 35 92 63
 IGF Nord 223
IGF Nord 223 04 34 35 92 78
04 34 35 92 78
 IGF Nord 223
IGF Nord 223 04 34 35 92 78
04 34 35 92 78
 IGF Nord 209b
IGF Nord 209b 04 34 35 92 89
04 34 35 92 89
 IGF Nord 215b
IGF Nord 215b 04 34 35 92 78
04 34 35 92 78
 IGF Nord 221a
IGF Nord 221a 04 34 35 93 07
04 34 35 93 07
 IGF Nord 223
IGF Nord 223 04 34 35 92 89
04 34 35 92 89