Team Julie PANNEQUIN
Signalization, plasticity and cancer
Project Monitoring circulating tumor DNA (ctDNA) to analyze colorectal cancer relapse
PRINCIPAL INVESTIGATOR

IGF staff involved
Julie PANNEQUIN
DR2, CNRS
Jean-Marc PASCUSSI
CRCN, Inserm
Caroline BONNANS
CRCN, Inserm
Jihane VITRE BOUBAKER
Thèse CNRS
Szimonetta HIDEG
AJT CDD, UM
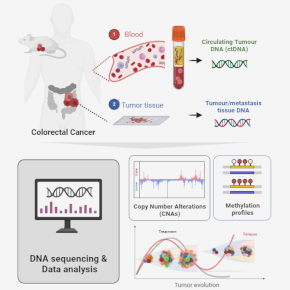
Our group is dedicated to the study of colorectal cancer (CRC) progression and recurrence by analyzing genomic sequencing data from clinical samples and preclinical models. We propose interdisciplinary research combining bioinformatics, experimental and translational research to develop new analytical methods to better understand the genomic dynamics of CRC and to improve the clinical follow-up of these tumors.
Tumor recurrence and metastasis are the leading causes of colorectal cancer (CRC) mortality, affecting nearly half of patients within 5-y after curative-intent therapy. Relapsed CRC may initially go undetected by standard surveillance, highlighting the need for enhanced monitoring to identify patients with post-treatment minimal residual disease (MRD). Circulating tumor DNA (ctDNA) measured by liquid biopsies is a promising non-invasive approach for continuous follow-up of patients with higher sensitivity than conventional imaging, enabling better detection of tumor dormancy, relapse and therapeutic resistance. Yet, its analysis is extremely challenging due to low fraction of tumor-derived cell-free DNA, and consensus on its clinical validity in CRC is currently lacking. Despite advances, current gold-standard next-generation sequencing (NGS) still shows limited sensitivity, especially in post-treatment remission. Additionally, it is resource-intensive and not always clinically accessible.
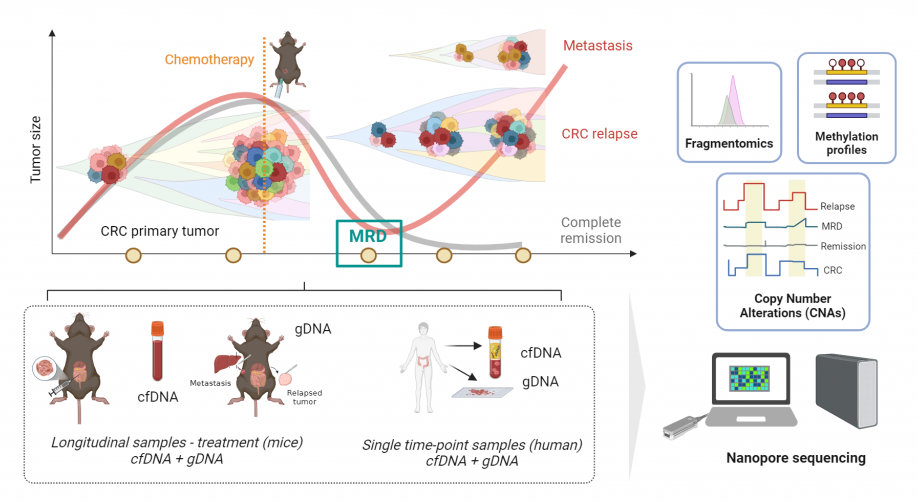
Here, our main goal is to improve the detection of CRC progression and recurrence by developing a minimally invasive, personalized and cost-effective monitoring strategy based on multimodal ctDNA analysis from a single blood sample. By exploiting the potential of third-generation nanopore-based sequencing, our approach aims to increase the sensitivity of ctDNA detection, especially when present at minimal levels after treatment (MRD), which can be considered a marker of early recurrence and therapeutic resistance.
To achieve this goal, the project includes:
- The use of mouse models for modelling clinical patterns of tumor progression and therapeutic resistance
- The use of CRC samples from patient plasma and tissue biopsies
- The development of computational methods for multimodal analysis of ctDNA from nanopore sequencing data
Main publications
• Pedrola A et al. (2022) Bioinformatics. 38(8):2374-2376.
• Yngvadottir B, Andreou A, Bassaganyas L, et al. (2022) Hum Mol Genet. 31(17):3001-3011
• Franch-Expósito S, Bassaganyas L, et al. (2020) Elife. 9:e50267
• Bassaganyas L, Pinyol R, Esteban-Fabró R, et al. (2020). Clin Cancer Res 26(23):6350-6361
• Pinyol R, Montal R, Bassaganyas L, et al. (2019) Gut. 68(6):1065-1075
• Bassaganyas L, Freedman G, et al. (2018) Hum Mutat. 39(1):167-171
• Bassaganyas L et al. (2013) Leukemia. 27(12):2376-2379
• Escaramís G, Tornador C, Bassaganyas L, et al. (2013) PLoS One. 8(5):e63377.
• Puente XS, et al. (2011) Nature. 475(7354):101-105


