This study was performed within the framework of the S.M.A.R.T. consortium whose goal is to design innovative tools for the detection and quantification of RNA modifications for clinical purpose (Région Occitanie and FEDER funding).
This project involves a cancer biology team (A. David, IGF), a bioinformatics team (E. Rivals, LIRMM) and a clinical mass spectrometry team (C. Hirtz, IRMB).
Chemical modifications of RNA, a.k.a. “epitranscriptomics”, represents another layer of gene expression control. Hitherto, more than 150 chemical marks have been discovered, from various organisms or RNA species, a complex, dynamic code, undetectable by conventional sequencing techniques. This study allowed us to identify a chemical mark whose presence on mRNA can promote cancer cell adaptation to conventional treatments. This mark is enriched in “cancer stem cells” (CSC) that fuel tumor growth. The involvement of epitranscriptomics in cancer evolution opens exciting perspectives for disease diagnosis and treatment.
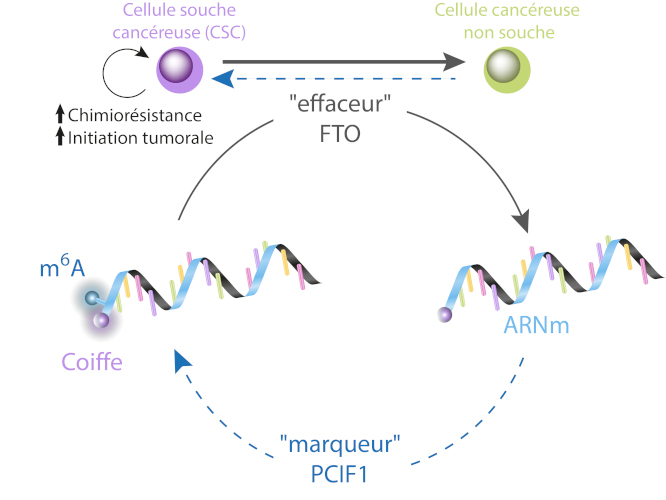
FTO targets « m6A » marks adjacent to the “cap” of messenger RNA (mRNA), genetic information-carrying intermediates. In patient derived cell lines, low FTO expression increases these m6A marks.
This, in turn, promotes the acquisition of “cancer stem cell” phenotype without modifying cell’s transcriptome.
Inhibiting PCIF1/PACAM, the nuclear “writer”, restores the sensitivity of cancer cell to conventional treatment.


